Escherichia coli
TAXONOMY
Domain: Bacteria;
Phylum: Pseudomonadota (Proteobacteria)
Class: Gammaproteobacteria
Order: Enterobacterales
Family: Enterobacteriaceae
Genus: Escherichia
MORPHOLOGY
Shape: Gram-negative rods (bacilli), typically 1–2 µm long
Gram Status: Gram-negative
Motility: Many strains are motile via peritrichous flagella; +P and 1‑type fimbriae aid attachment
Spore Formation: Non-spore-forming
Biofilm Formation: Strong biofilm producers on surfaces; fimbriae and surface factors play key roles
Other Traits: Lipopolysaccharide (LPS with O antigen), various K (capsular) antigens, and H (flagellar) antigens
NOTABLE TRAITS
Environmental/Food Safety: Widely found in intestines of warm-blooded animals; indicator of fecal contamination; thrives in varied environments Wikipedia
Virulence Factors:
Fimbriae and pili: type 1 fimbriae, P-pili (uropathogenic), F1C, S-pili
Toxins: Shiga toxins (Stx1/2) in EHEC/STEC, heat-labile/stable toxins in ETEC, hemolysin in UPEC
Iron acquisition & others: siderophores, adhesins, flagella
Survival Tolerance: Adapts to various pH, temperature; survives in food/water environments
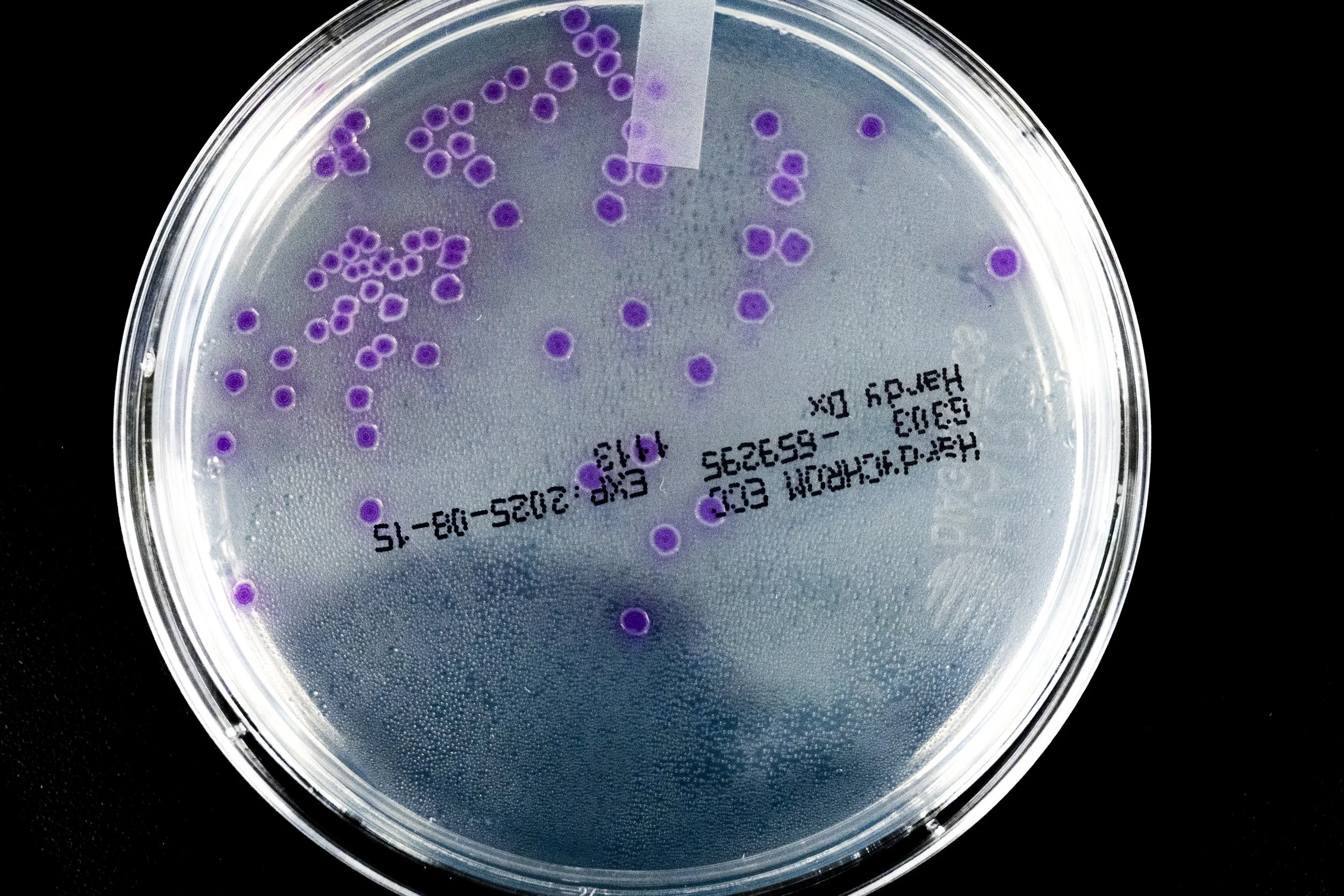



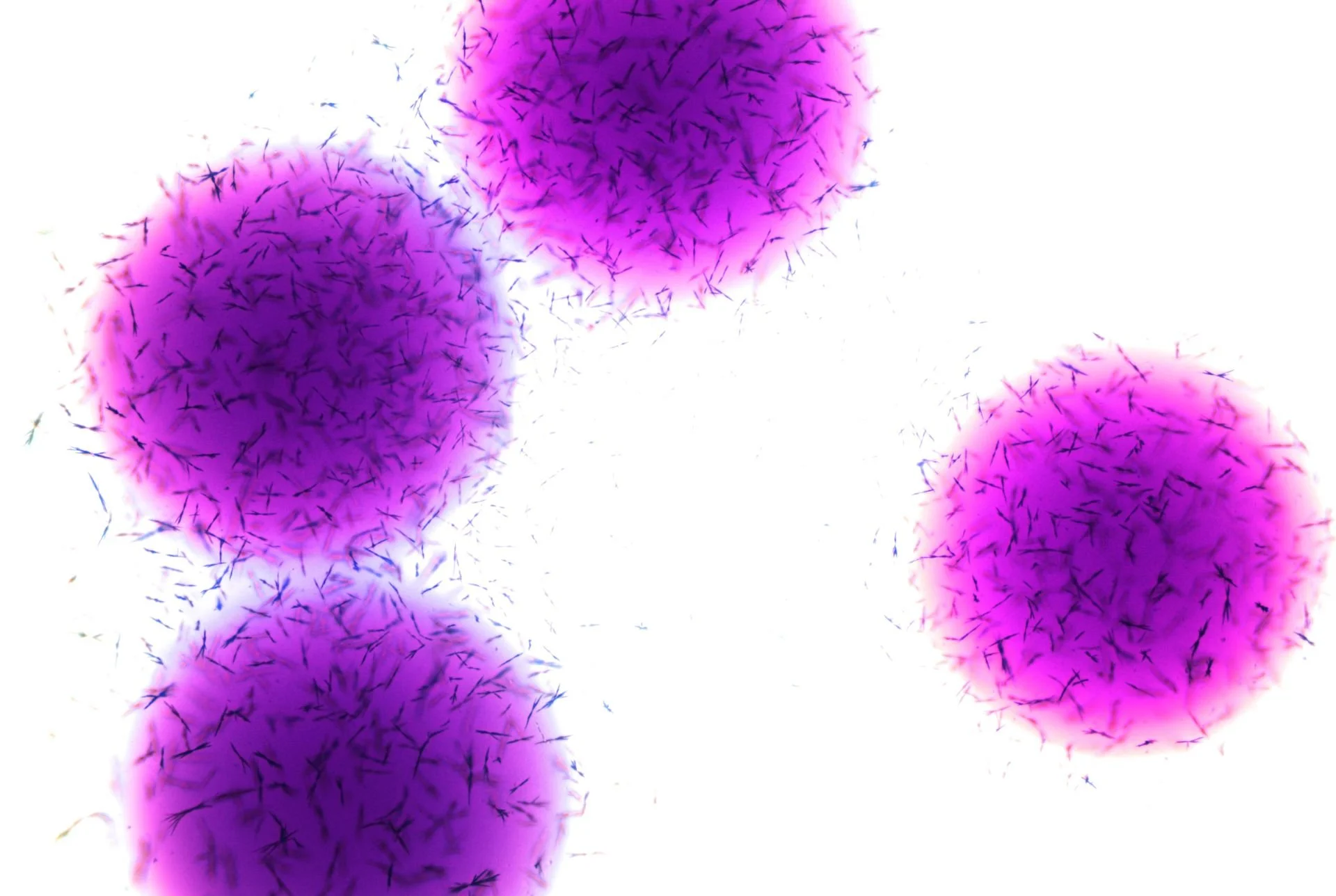
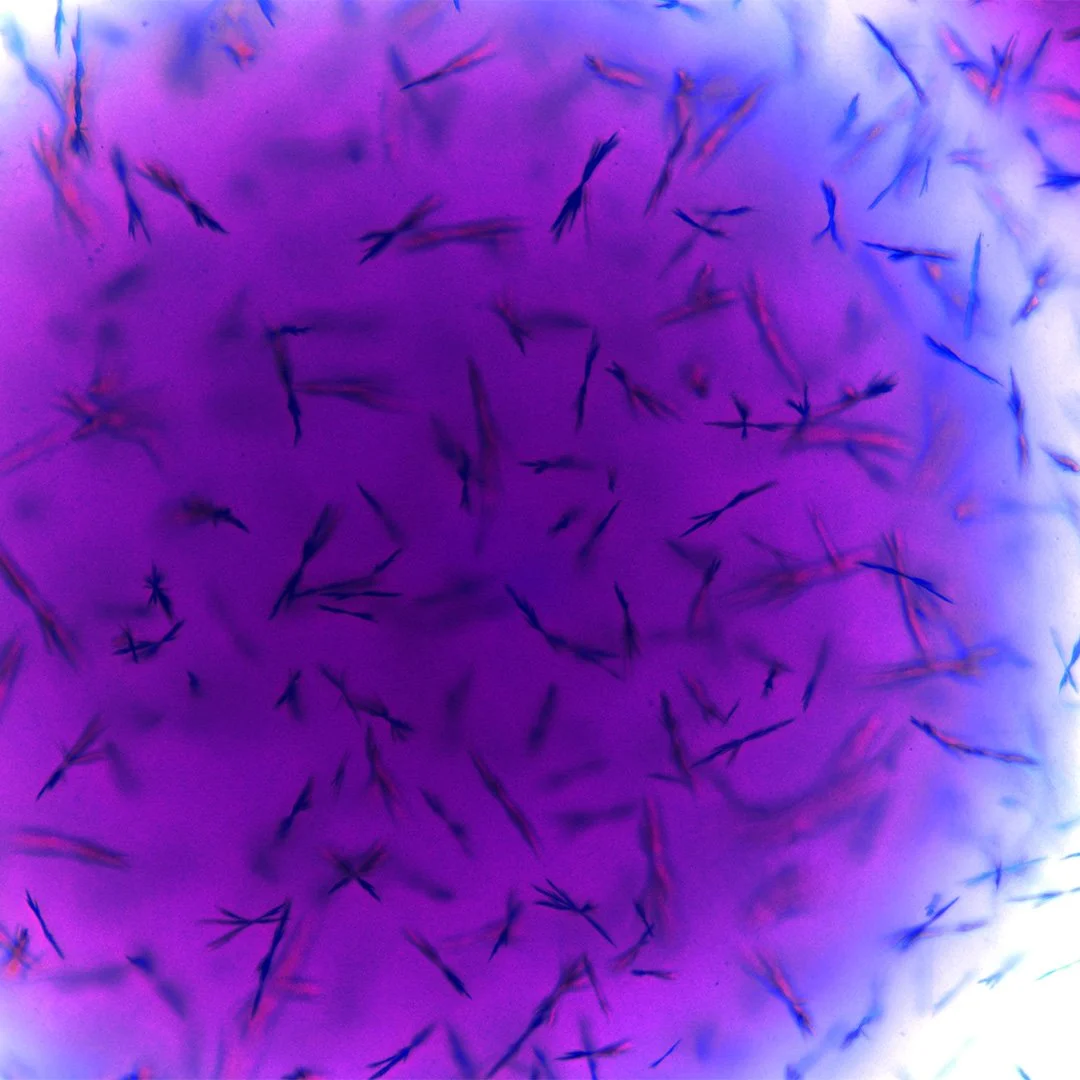
Seen through a microscope, Escherichia coli appears almost unassuming—short Gram-negative rods drifting through the field of view, some propelled by delicate peritrichous flagella that spin like microscopic propellers. Yet this modest organism has become one of the most studied life forms in the history of science.
The story of E. coli begins in 1885, when German pediatrician and bacteriologist Theodor Escherich first isolated the bacterium from infant fecal samples while investigating the microbial communities of the human gut. Escherich recognized that these microbes were not merely contaminants but permanent residents of the intestinal tract. His work helped establish the idea that humans coexist with vast microbial populations—a concept that today underpins our understanding of the microbiome.
For decades, E. coli served microbiologists as a reliable laboratory organism. It grows quickly, adapts easily to culture conditions, and its genetic systems proved remarkably accessible to experimentation. By the mid-20th century, it had become the workhorse of molecular biology, helping scientists decode the fundamental mechanisms of DNA replication, gene regulation, and protein synthesis.
One of the most transformative discoveries came through the work of François Jacob and Jacques Monod, who used E. coli to uncover how genes can be turned on and off through regulatory systems such as the lac operon. Their insights revealed how cells respond dynamically to their environment and earned them the 1965 Nobel Prize in Physiology or Medicine.
Yet E. coli also carries a more complex story. While most strains live harmlessly in the intestines of humans and animals, certain variants have acquired virulence factors that allow them to cause disease. Outbreaks linked to food contamination have periodically drawn global attention, highlighting the bacterium’s ability to evolve new capabilities through plasmids, toxins, and genetic exchange.
Despite these challenges, E. coli remains one of humanity’s most valuable scientific allies. It was among the first organisms used in recombinant DNA technology, enabling scientists to produce human insulin and other lifesaving medicines. Today, engineered strains are used to manufacture pharmaceuticals, study genetics, explore synthetic biology, and even develop new bio-based materials.
What began as a bacterium discovered in the human intestine has become a central figure in modern biology. Beneath the microscope, its simple rod-shaped form hides a remarkable story—one that links the human microbiome, the birth of molecular genetics, and the continuing promise of biotechnology.
In studying Escherichia coli, we are reminded that even the smallest organisms can illuminate the deepest principles of life—and that the tools of tomorrow’s science may already be quietly growing in a petri dish today.
